White Paper: Imaging-Based Viability Assessment of Cytostatic vs. Cytotoxic Chemotherapies in 3D Culture


Summary
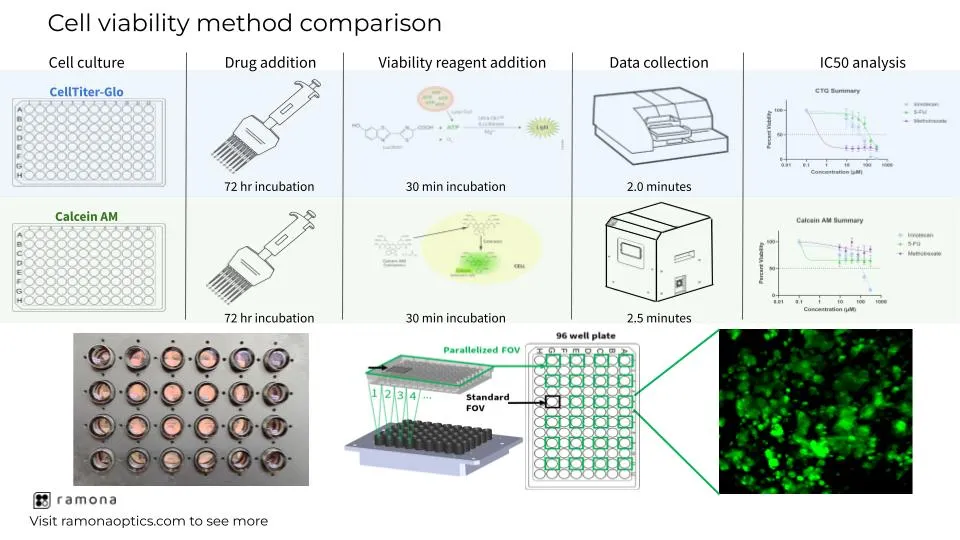
CellTiter-Glo (CTG) is the gold standard for assessment of cellular viability due to its ease of use and fast turnaround time for data analysis. Multiple commercially available systems are capable of assessing imaging-based viability, but the speed of acquisition and time-consuming analysis limits the sample throughput compared to CTG. Additionally, without an automated work cell to feed plates back and forth between the incubator and the imaging system, a researcher must spend a significant amount of hands-on time to complete the workflow. The Multi-Camera Array Microscope (MCAM™) by Ramona Optics offers unprecedented speed with its high-throughput imaging technology and cutting-edge data analysis tools. The MCAM™ can compete with the speed of acquisition and data analysis of a plate reader but offers a higher content of data gained by imaging. Imaging-based viability assessment offers the ability to collect data on a cellular level as opposed to the per-well data offered from plate readers. Cellular-level data provides information around heterogeneity of cellular toxicity responses to various treatment conditions, which is of great significance to in vitro heterogeneous disease models, such as cancer. Additionally, an imaging-based approach enables researchers to distinguish between drug cytotoxic and cytostatic mechanisms of action which is not possible with traditional CTG assays.
Here, we describe a viability-based workflow that offers a 10-fold decrease in processing time compared to traditional imaging-based viability assessment and a more accurate workflow compared to CTG. The MCAM™ is integrated with multiple commercially-available scheduling softwares for an entirely automated workflow, which further decreases the amount of hands-on time necessary. The purpose of this study is to demonstrate the increased efficiency in collecting and analyzing cell viability data offered by the MCAM™ technology as well as the information that is gained with an imaging-based viability approach compared to CTG. To accomplish this, a well-characterized colorectal cancer cell line, SW620, and various chemotherapeutic compounds that demonstrate cytostatic and cytotoxic phenotypes were used to compare imaging-based Calcein AM fluorescence to luminescence-based CTG readouts. This increase in imaging speed, data analytics, and phenotypic readouts can revolutionize the field of drug discovery and toxicology by drastically decreasing the time between image acquisition and data analysis and allowing for high-throughput screening capabilities.
Introduction
The CellTiter-Glo (CTG) luminescent cell viability assay by Promega (WI, USA) lyses the cell membrane within each well to release total cellular ATP. Using a combination of luciferin and luciferase, the reaction uses total cellular ATP released to induce a bioluminescent reaction that can be measured using a standard plate reader. This bioluminescent assay can be performed in a variety of well plate assays and yields a single value that represents the relative abundance of ATP per well that can be compared to control levels within the same plate.
Calcein AM is another widely used assay to determine cell viability. Unlike CellTiter-Glo, Calcein AM does not require cell lysis. Instead, Calcein AM can freely pass through the cell membrane where it undergoes cleavage by cytosolic esterases. The cytosolic esterase reaction results in a green fluorescent byproduct that is no longer membrane permeable and is retained by the cells. This reaction can then be quantified using a plate reader or by fluorescence microscopy. Plate readers are widely used for analyzing cellular viability data due to their speed of acquisition and ease of use. A caveat of using plate readers for either assay is that a single value is produced to represent hundreds to thousands of cells. Imaging-based approaches offer more granular information into the health of the individual cells in each well and can aid in deciphering between cytotoxic and cytostatic mechanisms of drug activity by maintaining cellular morphology data. Many researchers do not opt for imaging-based approaches to cell viability due to the increased time necessary for traditional microscopes to acquire data compared to plate readers as well as inadequate tools available for rapid analysis of the imaging data.
Additionally, the cost of automation-compatible, high-content microscopes and subsequent software that performs imaging-based data analysis can range from $500,000 — $1,000,000. Here, we describe the workflow of a new cell-based imaging platform, called the multi-camera array microscope (MCAM™), that can provide imaging-based cell viability assessment at comparable speed as a plate reader. We also directly compare the data output from the MCAM™ with the widely used CTG assay and provide insights into the conclusions that can be made using each technique.
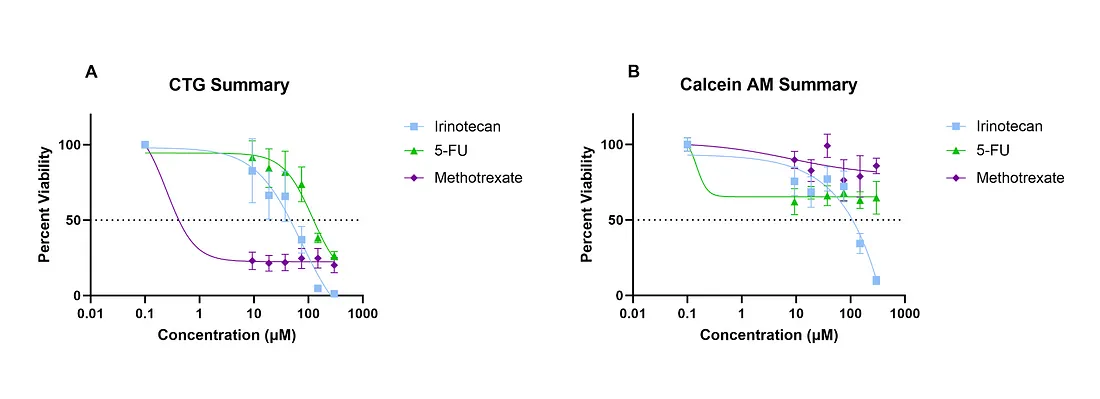
SW620 cells were grown in 3D suspension using 5 μL Matrigel domes plated in a 96 well black walled, clear bottom plate. 48 hours after plating SW620 cells in Matrigel domes, cells were treated with Irinotecan, 5-FU, or Methotrexate at concentrations ranging from 9 μM — 300 μM for 72hours. After 72 hours, the media was removed and replaced with 2 μM of Calcein AM and incubated at 37°C for 30 minutes. Calcein AM Fluorescence signal was measured with the MCAM™ by Ramona Optics by collecting 35–30 μm z-slices from each well. Total imaging time was 130 seconds for both channels across the entire 96 well plate. A maximum intensity projection was created and the total pixel area of Calcein AM signal was calculated for each well and normalized to the DMSO control group to generate the above dose response curve (B). After imaging was complete, CTG was added to each well and incubated for 30 minutes. Luminescence was measured using a plate reader and the viability was calculated and normalized to the DMSO control to generate the above CTG dose response curve (A). The scan time of the plate reader was 1 minute and 54 seconds.SW620 cells were grown in 3D suspension using 5 μL Matrigel domes plated in a 96 well black walled, clear bottom plate. 48 hours after plating SW620 cells in Matrigel domes, cells were treated with Irinotecan, 5-FU, or Methotrexate at concentrations ranging from 9 μM — 300 μM for 72hours. After 72 hours, the media was removed and replaced with 2 μM of Calcein AM and incubated at 37°C for 30 minutes. Calcein AM Fluorescence signal was measured with the MCAM™ by Ramona Optics by collecting 35–30 μm z-slices from each well. Total imaging time was 130 seconds for both channels across the entire 96 well plate. A maximum intensity projection was created and the total pixel area of Calcein AM signal was calculated for each well and normalized to the DMSO control group to generate the above dose response curve (B). After imaging was complete, CTG was added to each well and incubated for 30 minutes. Luminescence was measured using a plate reader and the viability was calculated and normalized to the DMSO control to generate the above CTG dose response curve (A). The scan time of the plate reader was 1 minute and 54 seconds.
Results
CellTiter-Glo versus Calcein AM dose response curves.
To better understand the impact different cell viability methods have on the overall conclusion of drug efficacy, we tested multiple well-established chemotherapeutic compounds against a colorectal cancer cell line, SW620, grown in 3D Matrigel domes. SW620 domes were drug treated for 72 hours and then treated with 2 μM of Calcein AM for 30 minutes prior to imaging with the MCAM™ 30 μm z-slices were collected over a 1 mm range. Total acquisition time was 130 seconds for both brightfield and green fluorescence channels. Images were analyzed using proprietary software by Ramona Optics that estimated the total number of green pixels per well (viable cell surface area). Once the images were collected, CTG was added to each well and incubated with agitation for 30 minutes to induce cell lysis. Total ATP was measured using a CLARIOstar plate reader (BMG LabTech, NC, USA). Percent viability was assessed for Calcein AM and CTG by normalizing the fluorescence or luminescence signal by the DMSO control. Dose response curves were generated and IC50 values calculated using GraphPad Prism software (Figure 1). Of the three chemotherapeutic compounds tested between each method, only Irinotecan yielded a similar IC50 value between CTG (74 μM) and Calcein AM (94 μM). The IC50 values for 5-Fluorouracil (5-FU) and Methotrexate were 119 μM and 0.25 μM respectively using the CTG method while the Calcein AM method did not display a viability lower than 50% for either compound (Figure 1), thus making an IC50 value impossible to generate. This discrepancy in drug efficacy led to a further analysis into the biology of these samples to determine which analysis method is more accurate.
Comparing brightfield and Calcein AM images in SW620 spheroids under different chemotherapeutics.
To verify whether the CTG or Calcein AM response curve best represents the biology, brightfield images were compared between DMSO control groups, 300 μM 5-FU, 300 μM Irinotecan, and 300 μM Methotrexate (Figure 2A). In brightfield, live cells appear round with well-defined membranes and bright centers. The cell morphology of dead cells is often distinctly different from live cells. Non-viable cells often have poorly defined cell membranes and appear darker and more condensed. The SW620 spheroids display characteristics of viable cells in DMSO control wells. Conversely, the SW620 spheroids treated with 300 μM of Irinotecan are smaller, more condensed, with poorly defined membranes. This aligns with the data from both the CTG and Calcein AM response curves which each display low viability in the 300 μM Irinotecan treatment group. The Methotrexate and 5-FU treatment groups show the largest discrepancy between the CTG and Calcein AM response curves (Figure 1) and average percent viability (Figure 2B). When viewing the brightfield images of the 300 μM Methotrexate treatment group, the SW620 spheroids appear rounded, bright, and viable, like the DMSO control group. To investigate further, the fluorescence intensity of Calcein AM stained samples were compared alongside the brightfield images (Figure 2B). In the top right corner of each panel in Figure 2B, the average percent viability for CTG (yellow) and Calcein AM signal (green) is displayed to compare to the actual images. Based on the brightfield morphology and the Calcein AM signal present in each treatment group, it is apparent that the CTG value is under-representing the viability of 5-FU and Methotrexate treatment groups.
Viability readout comparisons of cytotoxic, cytostatic, and resistant responses to chemotherapeutic treatment
Methotrexate, 5-FU, and Irinotecan are all classified as chemotherapeutics used to treat cancer, but each compound is involved in a different mechanism of action to inhibit growth or induce cell death. Methotrexate and 5-FU are anti-metabolites that inhibit DNA synthesis and subsequent cell division, while Irinotecan blocks DNA repair mechanisms, leading to apoptosis (1–3). It has been documented that assays that measure ATP content to determine cell viability can fail to distinguish between anti- proliferative/cytostatic and cytotoxic drug responses (4). In an age where 90% of new potential drug candidates fail in clinical trials, using phenotypic screening assays that can accurately characterize a variety of mechanisms of action is imperative (5). When comparing the two models of viability, Calcein AM fluorescence and CTG, the models only show a strong correlation between both measurements in the Irinotecan treatment group (Figure 3). The r2 value for the Irinotecan treatment group was 0.85 while the r2 values for Methotrexate and Irinotecan were 0.32 and 0.24, respectively. Exposure to Irinotecan leads to cell death while exposure to 5-FU and Methotrexate causes a reduction in cell division, which is apparent by the smaller spheroids formed in those treatment groups compared to the DMSO control (Figure 2). Additionally, resistance to 5-FU treatment has been documented in SW620 cells previously, which can also explain the diminished response (2) . Collectively, these data suggest that an imaging- based approach to viability assessment increases the accuracy of drug screening data by delineating antiproliferative activity from acute toxicity.
Conclusions
Imaging-based high content screening has become increasingly popular as a tool in the drug discovery field due to its ability to provide in depth information for complex biological systems. With over 90% of new compounds in development failing in clinical trials, researchers have migrated from 2D to 3D cell culture models to more accurately recapitulate human disease models. Dyes, probes, and genetic reporters for imaging have progressed significantly to address this need, but imaging platforms and imaging analysis workflows are slow and tedious due to the increase in imaging depth and content required for 3D cell culture.
Imaging and subsequent analysis has become the main bottleneck in many cell biology workflows, with imaging a 96 well plate in multiple z-planes taking between 30 and 40 minutes per plate, causing researchers to use traditional plate reader assays to accommodate their required throughput (6). Plate reader assays such as the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H- tetrazolium bromide (MTT) and CellTiter-Glo (CTG) condense information from thousands of cells and 3D organoids in each well into a single averaged value in under 2 minutes, but fail to provide information on the mechanism of action for the compound of interest.
Here, we present an overview of a workflow that can provide fast imaging-based well plate scanning and subsequent data analysis at the equivalent speed of a traditional CTG plate reader assay. The Calcein AM imaging of SW620 spheroids in Matrigel consisted of one field of view (FOV) per 96 well, 35 z-stacks per FOV, and 30 μm increments between each slice (Figure 1 and 2). The images were collected in approximately 80 seconds using the MCAM™ by Ramona Optics. The subsequent data analysis of the Calcein AM signal surface area was performed using proprietary software developed by Ramona Optics which processed the dataset in 70 seconds. All together, imaging and viability surface area assessment per well took around 2.5 minutes. The equivalent imaging parameters conducted on a Molecular Devices high-throughput imaging system would take between 30 and 40 minutes (6). These same plates were processed for CTG luminescence analysis using a BMG LabTech plate reader. The total time for CTG assessment and data output was two minutes. This demonstrates the 10-fold increase in speed obtained by using the MCAM™ compared to other imaging platforms as well as its speed compared to a plate reader based assay.
Additionally, we demonstrate the ability to distinguish between cytotoxic and cytostatic chemotherapeutic responses with the imaging-based approach which is not possible with other methods. CTG from the same assay plate revealed an overestimation of the toxicity of cytostatic compounds that inhibit cell proliferation. In the context of drug discovery and toxicology, the MCAM™ by Ramona Optics offers a cost effective, automation compatible, and high throughput method to perform imaging-based screening.
Methods
Cell Culture
SW620 cells were maintained in a 2D monolayer in 10 cm tissue culture treated dishes containing Phenol-Free DMEM with 10% FBS and 1% penicillin/streptomycin. For the viability assay, SW620 cells treated with Trypsin for 5 minutes and resuspended in Matrigel at a concentration of 500,000 cells/mL. 5 μL SW620 Matrigel domes were plated in a clear bottom, black walled 96 well plate. After Matrigel solidification, 100 μL of Phenol-Free DMEM with 10% FBS and 1% penicillin/streptomycin was added to each well. SW620 cells embedded in Matrigel were grown for 72 hours to allow for spheroid formation prior to chemotherapeutic treatment. The experiments described in this white paper were conducted in biological duplicate and technical triplicate.
Chemotherapeutic preparation and addition
Lyophilized Irinotecan, Methotrexate, and 5-Fluorouracil were purchased from Cayman Chemical (MI, USA). Each compound was resuspended in DMSO at the manufacturer’s recommended solubility. A 6 point serial dilution of Irinotecan, Methotrexate, and 5-Fluorouracil was prepared at 300 μM, 150 μM, 75 μM, 37.5 μM, 18.75 μM, and 9.38 μM. 6 wells were left as a 1% DMSO control for untreated comparisons. Each compound was diluted to the specified concentration in Phenol- Free Complete DMEM media prior to addition to the SW620 cells embedded in Matrigel domes, which were treated for 72 hours prior to viability assessment.
Calcein AM Imaging using the Multi-Camera Array Microscope (MCAM™)
After 72 hours of incubation with chemotherapeutics, media was removed from each well and replaced with a 2 μM solution of Calcein AM purchased from Biotium (CA, USA). Cells were then incubated with the Calcein AM solution at 37 °C for 30 minutes. A full 96 well plate scan was performed using the MCAM™ by Ramona Optics in brightfield and green fluorescence. Each scan consisted of 35 - 30 μm z-slices and took 90 seconds to complete.
CellTiter-Glo Analysis
After Calcein AM fluorescence imaging was completed, the 2 μM solution of Calcein AM was removed from each well and replaced with 100 μL of 3D CellTiter-Glo from Promega. Plates were incubated at room temperature with gentle agitation for 30 minutes prior to luminescence reading on a CLARIOstar Plus plate reader by BMG Labtech, which acquired relative luminescence values for the entire plate in 1 minute and 54 seconds.
Data Analysis
For the imaging-based approach, the 35 z-slices collected during the Calcein AM fluorescence scan were compiled into a maximum intensity projection. The total number of fluorescent pixels were calculated from the maximum intensity projections of each well to provide the total viable surface area. The total viable surface area from each well was normalized to the 1% DMSO control to calculate a percent viability for each treatment group. For the CellTiter-Glo data analysis, the RLU values generated from the plate reader were normalized to the 1% DMSO control to calculate the percent viability for each treatment group. All data points for Calcein AM fluorescence and CTG based luminescence were compiled in GraphPad Prism and a nonlinear regression curve analysis was performed.
References
- Koźmiński, Przemysław et al. “Overview of Dual-Acting Drug Methotrexate in Different Neurological Diseases, Autoimmune Pathologies and Cancers.” International journal of molecular sciences vol. 21,10 3483. 14 May. 2020, doi:10.3390/ijms21103483
- Blondy, Sabrina et al. “5-Fluorouracil resistance mechanisms in colorectal cancer: From classical pathways to promising processes.” Cancer science vol. 111,9 (2020): 3142–3154. doi:10.1111/cas.14532
- Gilbert, D C et al. “Topoisomerase I inhibition in colorectal cancer: biomarkers and therapeutic targets.” British journal of cancer vol. 106,1 (2012): 18–24. doi:10.1038/bjc.2011.498
- Chan, Grace Ka Yan et al. “A simple high-content cell cycle assay reveals frequentdiscrepancies between cell number and ATP and MTS proliferation assays.” PloS one vol. 8,5 e63583. 17 May. 2013, doi:10.1371/journal.pone.0063583
- Zheng, Wei et al. “Phenotypic screens as a renewed approach for drug discovery.” Drug discovery today vol. 18,21–22 (2013): 1067–73. doi:10.1016/j.drudis.2013.07.001
- Molecular Devices. Retrieved January 4th, 2024, from https://www.moleculardevices.com/products/cellular-imaging-systems

White Paper: Imaging-Based Viability Assessment of Cytostatic vs. Cytotoxic Chemotherapies in 3D Culture
Summary
CellTiter-Glo (CTG) is the gold standard for assessment of cellular viability due to its ease of use and fast turnaround time for data analysis. Multiple commercially available systems are capable of assessing imaging-based viability, but the speed of acquisition and time-consuming analysis limits the sample throughput compared to CTG. Additionally, without an automated work cell to feed plates back and forth between the incubator and the imaging system, a researcher must spend a significant amount of hands-on time to complete the workflow. The Multi-Camera Array Microscope (MCAM™) by Ramona Optics offers unprecedented speed with its high-throughput imaging technology and cutting-edge data analysis tools. The MCAM™ can compete with the speed of acquisition and data analysis of a plate reader but offers a higher content of data gained by imaging. Imaging-based viability assessment offers the ability to collect data on a cellular level as opposed to the per-well data offered from plate readers. Cellular-level data provides information around heterogeneity of cellular toxicity responses to various treatment conditions, which is of great significance to in vitro heterogeneous disease models, such as cancer. Additionally, an imaging-based approach enables researchers to distinguish between drug cytotoxic and cytostatic mechanisms of action which is not possible with traditional CTG assays.
Here, we describe a viability-based workflow that offers a 10-fold decrease in processing time compared to traditional imaging-based viability assessment and a more accurate workflow compared to CTG. The MCAM™ is integrated with multiple commercially-available scheduling softwares for an entirely automated workflow, which further decreases the amount of hands-on time necessary. The purpose of this study is to demonstrate the increased efficiency in collecting and analyzing cell viability data offered by the MCAM™ technology as well as the information that is gained with an imaging-based viability approach compared to CTG. To accomplish this, a well-characterized colorectal cancer cell line, SW620, and various chemotherapeutic compounds that demonstrate cytostatic and cytotoxic phenotypes were used to compare imaging-based Calcein AM fluorescence to luminescence-based CTG readouts. This increase in imaging speed, data analytics, and phenotypic readouts can revolutionize the field of drug discovery and toxicology by drastically decreasing the time between image acquisition and data analysis and allowing for high-throughput screening capabilities.

Introduction
The CellTiter-Glo (CTG) luminescent cell viability assay by Promega (WI, USA) lyses the cell membrane within each well to release total cellular ATP. Using a combination of luciferin and luciferase, the reaction uses total cellular ATP released to induce a bioluminescent reaction that can be measured using a standard plate reader. This bioluminescent assay can be performed in a variety of well plate assays and yields a single value that represents the relative abundance of ATP per well that can be compared to control levels within the same plate.
Calcein AM is another widely used assay to determine cell viability. Unlike CellTiter-Glo, Calcein AM does not require cell lysis. Instead, Calcein AM can freely pass through the cell membrane where it undergoes cleavage by cytosolic esterases. The cytosolic esterase reaction results in a green fluorescent byproduct that is no longer membrane permeable and is retained by the cells. This reaction can then be quantified using a plate reader or by fluorescence microscopy. Plate readers are widely used for analyzing cellular viability data due to their speed of acquisition and ease of use. A caveat of using plate readers for either assay is that a single value is produced to represent hundreds to thousands of cells. Imaging-based approaches offer more granular information into the health of the individual cells in each well and can aid in deciphering between cytotoxic and cytostatic mechanisms of drug activity by maintaining cellular morphology data. Many researchers do not opt for imaging-based approaches to cell viability due to the increased time necessary for traditional microscopes to acquire data compared to plate readers as well as inadequate tools available for rapid analysis of the imaging data.
Additionally, the cost of automation-compatible, high-content microscopes and subsequent software that performs imaging-based data analysis can range from $500,000 — $1,000,000. Here, we describe the workflow of a new cell-based imaging platform, called the multi-camera array microscope (MCAM™), that can provide imaging-based cell viability assessment at comparable speed as a plate reader. We also directly compare the data output from the MCAM™ with the widely used CTG assay and provide insights into the conclusions that can be made using each technique.

SW620 cells were grown in 3D suspension using 5 μL Matrigel domes plated in a 96 well black walled, clear bottom plate. 48 hours after plating SW620 cells in Matrigel domes, cells were treated with Irinotecan, 5-FU, or Methotrexate at concentrations ranging from 9 μM — 300 μM for 72hours. After 72 hours, the media was removed and replaced with 2 μM of Calcein AM and incubated at 37°C for 30 minutes. Calcein AM Fluorescence signal was measured with the MCAM™ by Ramona Optics by collecting 35–30 μm z-slices from each well. Total imaging time was 130 seconds for both channels across the entire 96 well plate. A maximum intensity projection was created and the total pixel area of Calcein AM signal was calculated for each well and normalized to the DMSO control group to generate the above dose response curve (B). After imaging was complete, CTG was added to each well and incubated for 30 minutes. Luminescence was measured using a plate reader and the viability was calculated and normalized to the DMSO control to generate the above CTG dose response curve (A). The scan time of the plate reader was 1 minute and 54 seconds.SW620 cells were grown in 3D suspension using 5 μL Matrigel domes plated in a 96 well black walled, clear bottom plate. 48 hours after plating SW620 cells in Matrigel domes, cells were treated with Irinotecan, 5-FU, or Methotrexate at concentrations ranging from 9 μM — 300 μM for 72hours. After 72 hours, the media was removed and replaced with 2 μM of Calcein AM and incubated at 37°C for 30 minutes. Calcein AM Fluorescence signal was measured with the MCAM™ by Ramona Optics by collecting 35–30 μm z-slices from each well. Total imaging time was 130 seconds for both channels across the entire 96 well plate. A maximum intensity projection was created and the total pixel area of Calcein AM signal was calculated for each well and normalized to the DMSO control group to generate the above dose response curve (B). After imaging was complete, CTG was added to each well and incubated for 30 minutes. Luminescence was measured using a plate reader and the viability was calculated and normalized to the DMSO control to generate the above CTG dose response curve (A). The scan time of the plate reader was 1 minute and 54 seconds.
Results
CellTiter-Glo versus Calcein AM dose response curves.
To better understand the impact different cell viability methods have on the overall conclusion of drug efficacy, we tested multiple well-established chemotherapeutic compounds against a colorectal cancer cell line, SW620, grown in 3D Matrigel domes. SW620 domes were drug treated for 72 hours and then treated with 2 μM of Calcein AM for 30 minutes prior to imaging with the MCAM™ 30 μm z-slices were collected over a 1 mm range. Total acquisition time was 130 seconds for both brightfield and green fluorescence channels. Images were analyzed using proprietary software by Ramona Optics that estimated the total number of green pixels per well (viable cell surface area). Once the images were collected, CTG was added to each well and incubated with agitation for 30 minutes to induce cell lysis. Total ATP was measured using a CLARIOstar plate reader (BMG LabTech, NC, USA). Percent viability was assessed for Calcein AM and CTG by normalizing the fluorescence or luminescence signal by the DMSO control. Dose response curves were generated and IC50 values calculated using GraphPad Prism software (Figure 1). Of the three chemotherapeutic compounds tested between each method, only Irinotecan yielded a similar IC50 value between CTG (74 μM) and Calcein AM (94 μM). The IC50 values for 5-Fluorouracil (5-FU) and Methotrexate were 119 μM and 0.25 μM respectively using the CTG method while the Calcein AM method did not display a viability lower than 50% for either compound (Figure 1), thus making an IC50 value impossible to generate. This discrepancy in drug efficacy led to a further analysis into the biology of these samples to determine which analysis method is more accurate.
Comparing brightfield and Calcein AM images in SW620 spheroids under different chemotherapeutics.
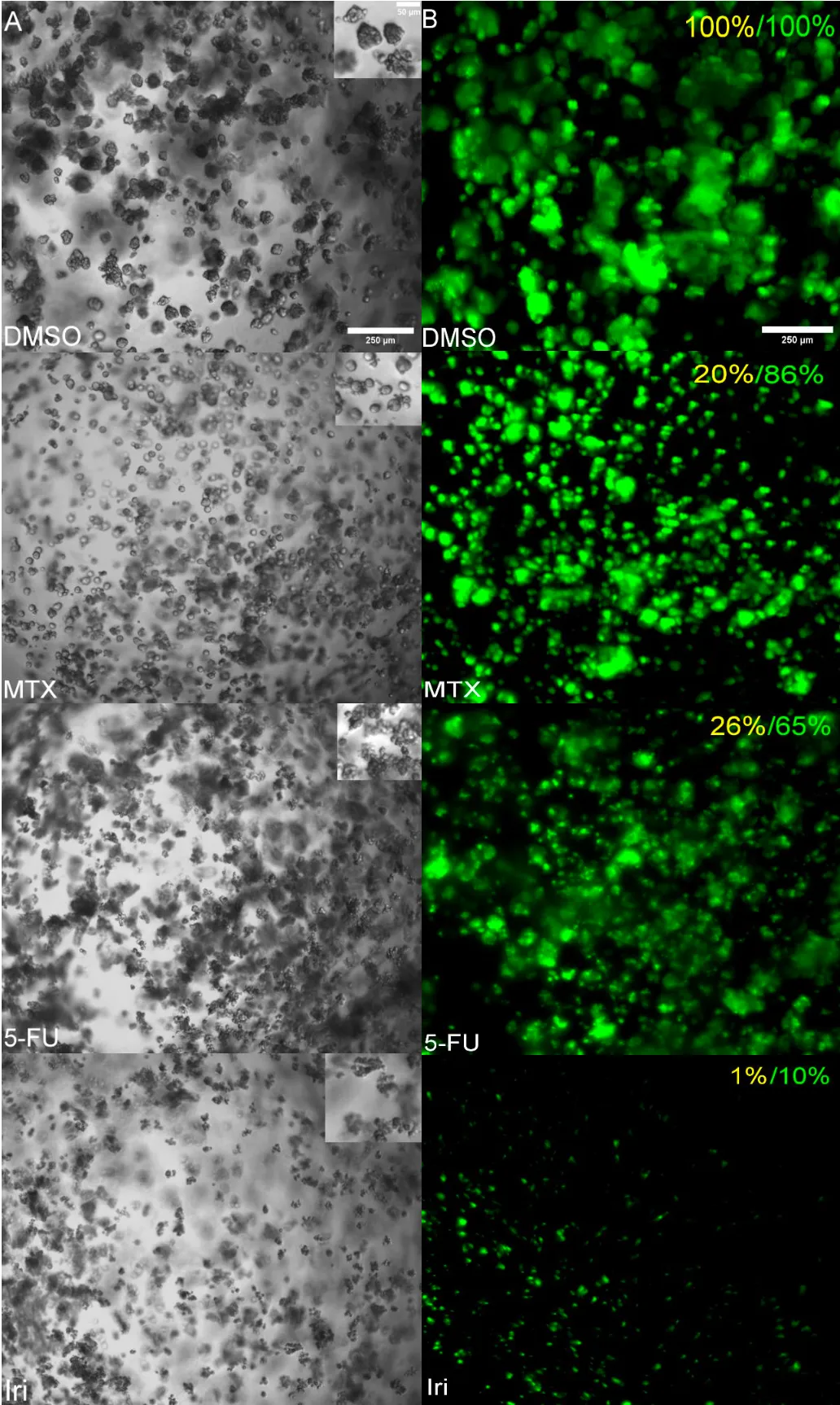
To verify whether the CTG or Calcein AM response curve best represents the biology, brightfield images were compared between DMSO control groups, 300 μM 5-FU, 300 μM Irinotecan, and 300 μM Methotrexate (Figure 2A). In brightfield, live cells appear round with well-defined membranes and bright centers. The cell morphology of dead cells is often distinctly different from live cells. Non-viable cells often have poorly defined cell membranes and appear darker and more condensed. The SW620 spheroids display characteristics of viable cells in DMSO control wells. Conversely, the SW620 spheroids treated with 300 μM of Irinotecan are smaller, more condensed, with poorly defined membranes. This aligns with the data from both the CTG and Calcein AM response curves which each display low viability in the 300 μM Irinotecan treatment group. The Methotrexate and 5-FU treatment groups show the largest discrepancy between the CTG and Calcein AM response curves (Figure 1) and average percent viability (Figure 2B). When viewing the brightfield images of the 300 μM Methotrexate treatment group, the SW620 spheroids appear rounded, bright, and viable, like the DMSO control group. To investigate further, the fluorescence intensity of Calcein AM stained samples were compared alongside the brightfield images (Figure 2B). In the top right corner of each panel in Figure 2B, the average percent viability for CTG (yellow) and Calcein AM signal (green) is displayed to compare to the actual images. Based on the brightfield morphology and the Calcein AM signal present in each treatment group, it is apparent that the CTG value is under-representing the viability of 5-FU and Methotrexate treatment groups.
Viability readout comparisons of cytotoxic, cytostatic, and resistant responses to chemotherapeutic treatment
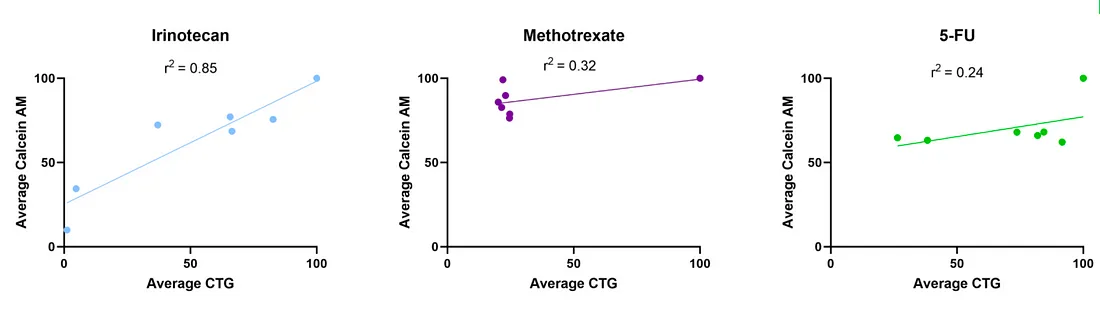
Methotrexate, 5-FU, and Irinotecan are all classified as chemotherapeutics used to treat cancer, but each compound is involved in a different mechanism of action to inhibit growth or induce cell death. Methotrexate and 5-FU are anti-metabolites that inhibit DNA synthesis and subsequent cell division, while Irinotecan blocks DNA repair mechanisms, leading to apoptosis (1–3). It has been documented that assays that measure ATP content to determine cell viability can fail to distinguish between anti- proliferative/cytostatic and cytotoxic drug responses (4). In an age where 90% of new potential drug candidates fail in clinical trials, using phenotypic screening assays that can accurately characterize a variety of mechanisms of action is imperative (5). When comparing the two models of viability, Calcein AM fluorescence and CTG, the models only show a strong correlation between both measurements in the Irinotecan treatment group (Figure 3). The r2 value for the Irinotecan treatment group was 0.85 while the r2 values for Methotrexate and Irinotecan were 0.32 and 0.24, respectively. Exposure to Irinotecan leads to cell death while exposure to 5-FU and Methotrexate causes a reduction in cell division, which is apparent by the smaller spheroids formed in those treatment groups compared to the DMSO control (Figure 2). Additionally, resistance to 5-FU treatment has been documented in SW620 cells previously, which can also explain the diminished response (2) . Collectively, these data suggest that an imaging- based approach to viability assessment increases the accuracy of drug screening data by delineating antiproliferative activity from acute toxicity.
Conclusions
Imaging-based high content screening has become increasingly popular as a tool in the drug discovery field due to its ability to provide in depth information for complex biological systems. With over 90% of new compounds in development failing in clinical trials, researchers have migrated from 2D to 3D cell culture models to more accurately recapitulate human disease models. Dyes, probes, and genetic reporters for imaging have progressed significantly to address this need, but imaging platforms and imaging analysis workflows are slow and tedious due to the increase in imaging depth and content required for 3D cell culture.


Imaging and subsequent analysis has become the main bottleneck in many cell biology workflows, with imaging a 96 well plate in multiple z-planes taking between 30 and 40 minutes per plate, causing researchers to use traditional plate reader assays to accommodate their required throughput (6). Plate reader assays such as the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H- tetrazolium bromide (MTT) and CellTiter-Glo (CTG) condense information from thousands of cells and 3D organoids in each well into a single averaged value in under 2 minutes, but fail to provide information on the mechanism of action for the compound of interest.
Here, we present an overview of a workflow that can provide fast imaging-based well plate scanning and subsequent data analysis at the equivalent speed of a traditional CTG plate reader assay. The Calcein AM imaging of SW620 spheroids in Matrigel consisted of one field of view (FOV) per 96 well, 35 z-stacks per FOV, and 30 μm increments between each slice (Figure 1 and 2). The images were collected in approximately 80 seconds using the MCAM™ by Ramona Optics. The subsequent data analysis of the Calcein AM signal surface area was performed using proprietary software developed by Ramona Optics which processed the dataset in 70 seconds. All together, imaging and viability surface area assessment per well took around 2.5 minutes. The equivalent imaging parameters conducted on a Molecular Devices high-throughput imaging system would take between 30 and 40 minutes (6). These same plates were processed for CTG luminescence analysis using a BMG LabTech plate reader. The total time for CTG assessment and data output was two minutes. This demonstrates the 10-fold increase in speed obtained by using the MCAM™ compared to other imaging platforms as well as its speed compared to a plate reader based assay.
Additionally, we demonstrate the ability to distinguish between cytotoxic and cytostatic chemotherapeutic responses with the imaging-based approach which is not possible with other methods. CTG from the same assay plate revealed an overestimation of the toxicity of cytostatic compounds that inhibit cell proliferation. In the context of drug discovery and toxicology, the MCAM™ by Ramona Optics offers a cost effective, automation compatible, and high throughput method to perform imaging-based screening.
Methods
Cell Culture
SW620 cells were maintained in a 2D monolayer in 10 cm tissue culture treated dishes containing Phenol-Free DMEM with 10% FBS and 1% penicillin/streptomycin. For the viability assay, SW620 cells treated with Trypsin for 5 minutes and resuspended in Matrigel at a concentration of 500,000 cells/mL. 5 μL SW620 Matrigel domes were plated in a clear bottom, black walled 96 well plate. After Matrigel solidification, 100 μL of Phenol-Free DMEM with 10% FBS and 1% penicillin/streptomycin was added to each well. SW620 cells embedded in Matrigel were grown for 72 hours to allow for spheroid formation prior to chemotherapeutic treatment. The experiments described in this white paper were conducted in biological duplicate and technical triplicate.
Chemotherapeutic preparation and addition
Lyophilized Irinotecan, Methotrexate, and 5-Fluorouracil were purchased from Cayman Chemical (MI, USA). Each compound was resuspended in DMSO at the manufacturer’s recommended solubility. A 6 point serial dilution of Irinotecan, Methotrexate, and 5-Fluorouracil was prepared at 300 μM, 150 μM, 75 μM, 37.5 μM, 18.75 μM, and 9.38 μM. 6 wells were left as a 1% DMSO control for untreated comparisons. Each compound was diluted to the specified concentration in Phenol- Free Complete DMEM media prior to addition to the SW620 cells embedded in Matrigel domes, which were treated for 72 hours prior to viability assessment.
Calcein AM Imaging using the Multi-Camera Array Microscope (MCAM™)
After 72 hours of incubation with chemotherapeutics, media was removed from each well and replaced with a 2 μM solution of Calcein AM purchased from Biotium (CA, USA). Cells were then incubated with the Calcein AM solution at 37 °C for 30 minutes. A full 96 well plate scan was performed using the MCAM™ by Ramona Optics in brightfield and green fluorescence. Each scan consisted of 35 - 30 μm z-slices and took 90 seconds to complete.
CellTiter-Glo Analysis
After Calcein AM fluorescence imaging was completed, the 2 μM solution of Calcein AM was removed from each well and replaced with 100 μL of 3D CellTiter-Glo from Promega. Plates were incubated at room temperature with gentle agitation for 30 minutes prior to luminescence reading on a CLARIOstar Plus plate reader by BMG Labtech, which acquired relative luminescence values for the entire plate in 1 minute and 54 seconds.
Data Analysis
For the imaging-based approach, the 35 z-slices collected during the Calcein AM fluorescence scan were compiled into a maximum intensity projection. The total number of fluorescent pixels were calculated from the maximum intensity projections of each well to provide the total viable surface area. The total viable surface area from each well was normalized to the 1% DMSO control to calculate a percent viability for each treatment group. For the CellTiter-Glo data analysis, the RLU values generated from the plate reader were normalized to the 1% DMSO control to calculate the percent viability for each treatment group. All data points for Calcein AM fluorescence and CTG based luminescence were compiled in GraphPad Prism and a nonlinear regression curve analysis was performed.
References
- Koźmiński, Przemysław et al. “Overview of Dual-Acting Drug Methotrexate in Different Neurological Diseases, Autoimmune Pathologies and Cancers.” International journal of molecular sciences vol. 21,10 3483. 14 May. 2020, doi:10.3390/ijms21103483
- Blondy, Sabrina et al. “5-Fluorouracil resistance mechanisms in colorectal cancer: From classical pathways to promising processes.” Cancer science vol. 111,9 (2020): 3142–3154. doi:10.1111/cas.14532
- Gilbert, D C et al. “Topoisomerase I inhibition in colorectal cancer: biomarkers and therapeutic targets.” British journal of cancer vol. 106,1 (2012): 18–24. doi:10.1038/bjc.2011.498
- Chan, Grace Ka Yan et al. “A simple high-content cell cycle assay reveals frequentdiscrepancies between cell number and ATP and MTS proliferation assays.” PloS one vol. 8,5 e63583. 17 May. 2013, doi:10.1371/journal.pone.0063583
- Zheng, Wei et al. “Phenotypic screens as a renewed approach for drug discovery.” Drug discovery today vol. 18,21–22 (2013): 1067–73. doi:10.1016/j.drudis.2013.07.001
- Molecular Devices. Retrieved January 4th, 2024, from https://www.moleculardevices.com/products/cellular-imaging-systems